Research
All eukaryotic cells are covered in a dense layer of carbohydrates that are key to cell-cell communication, development, neurobiology and immune function. In addition, many human viruses interact with cell-surface carbohydrates as receptors on their host cells in order to initiate infection. Carbohydrate structures are expressed in tissue- and species-specific patterns, and the availability of the receptor carbohydrate influences viral tissue tropism and host range. Viruses also encounter carbohydrates in the extracellular matrix and the environment on their way from one host cell to another. Some of these extracellular glycans sequester viruses from cells and are part of innate immunity, while others can render viruses more infectious. In any case, the specificity and affinity of viral proteins for carbohydrates determines viral spread, transmission and pathogenesis. However, protein-carbohydrate interactions are less well understood than protein-protein or protein-nucleic acid interactions. Moreover, virus-carbohydrate interactions are multivalent, with many shallow, low-affinity binding sites acting in concert.
Our aim is to determine the rules of engagement between viruses and relevant carbohydrate structures. We determine the specificity of viral proteins for carbohydrates by glycobiological and biophysics approaches and solve the structures of cognate protein-carbohydrate complexes by x-ray crystallography. In particular, we ask the following questions:
- By which molecular interactions does the protein recognize its carbohydrate ligand(s)?
- How do changes in these interactions impart binding specificity and strength?
- What spectrum of carbohydrates is bound by a specific protein?
- How are the biological functions of the protein correlated with its carbohydrate binding properties?
The answers to these questions advance our understanding of viral spread and infection and provide the structural basis for the design of novel antivirals. In addition, they shape our growing understanding of protein-carbohydrate interactions.
Our research is funded by the
- Emmy Noether Programme of the DFG
- SFB 765
- MPG
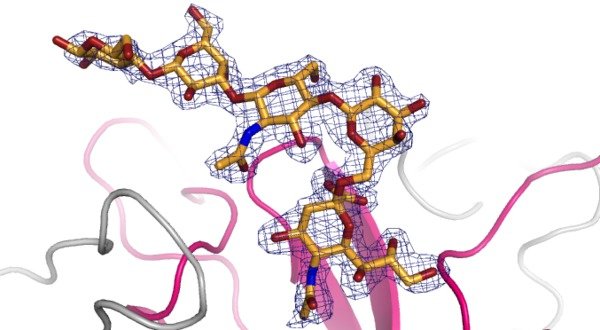
Structural information is key to understanding protein-carbohydrate recognition processes. Even with the glycan ligand and the structure of the binding protein known, it is challenging to predict the structure of the complex. Glycans are conformationally flexible and it is now known which conformation of the carbohydrate is bound. In many cases, the binding sites are shallow and hard to discriminate from unrelated protein surface by docking programs. Structure determination of protein-carbohydrate complexes is particularly synergistic with approaches that define the glycan binding specificity of the protein, such as glycan microarray screening and ligand-based NMR techniques. The binding specificity information can focus crystallographic efforts onto few carbohydrate structures of interest. Likewise, knowledge of the structure of the complex can rationalise glycan microarray data, e.g. by identifying amino acids that block interactions with a particular group of glycan motifs, or that provide favourable contacts to others.